Test repository for EEG project.
Data originally from: https://archive.ics.uci.edu/ml/datasets/eeg+database
Experiment description:
There were two groups of subjects: alcoholic and control.
Each subject was exposed to either a single stimulus (S1) or to two stimuli (S1 and S2) which were pictures of objects chosen from the 1980 Snodgrass and Vanderwart picture set. When two stimuli were shown, they were presented in either a matched condition where S1 was identical to S2 or in a non-matched condition where S1 differed from S2.
There were 122 subjects and each subject completed 120 trials where different stimuli were shown. The electrode positions were located at standard sites
Data download: https://archive.ics.uci.edu/ml/machine-learning-databases/eeg-mld/
There are three versions of the EEG data set.
- The Small Data Set
The small data set (smni97_eeg_data.tar.gz) contains data for the 2 subjects, alcoholic a_co2a0000364 and control c_co2c0000337. For each of the 3 matching paradigms, c_1 (one presentation only), c_m (match to previous presentation) and c_n (no-match to previous presentation), 10 runs are shown.
- The Large Data Set
The large data set (SMNI_CMI_TRAIN.tar.gz and SMNI_CMI_TEST.tar.gz) contains data for 10 alcoholic and 10 control subjects, with 10 runs (=trials) per subject per paradigm. Since there are three paradigms (one stimulus, two matched stimuli, two non-matched stimuli) there are 30 runs for each subjects. The test data used the same 10 alcoholic and 10 control subjects as with the training data, but with 10 out-of-sample runs per subject per paradigm.
3. The Full Data Set
This data set, eeg_full.tar contains all 120 trials for 122 subjects. The entire set of data is about 700 MBytes.
Actually, each subject has a certain number of trials missing (those were the trials that were unsuccessfull.) So each partcipant's folder contains a different number of .gz files. For example participant 365 has 93 trials on record, while participant 364 has 88 trials on record.
The dataset is on the UCI website at this link: https://archive.ics.uci.edu/ml/machine-learning-databases/eeg-mld/
NOTE: There are 17 trials with empty files in co2c1000367. Some trials have "err" notices, e.g., search/grep for "err" and see "S2 match err" or "S2 nomatch err" etc.
DATA DESCRIPTION
The folder SMNI_CMI_TRAIN contains the train data from the The Large Data Set (see description above). The dataset contains the data for 20 participants, 10 in the control group and 10 in the alcoholic group. Each participant identifies a subfolder, such as co2a0000364, co2a0000365 etc. If the subfolder is named co2a0000364, for example, that's participant "364", and we know that he or she was in the alcoholic group because of the "a" in "co2a..." If there was a "c" in place of the a, that would be a participant in the control group.
Each partcipant subfolder contains 30 .gz files (such as co2a0000364.rd.025.gz, co2a0000364.rd.031.gz) Each gz. file stands for one trial, and there are 10 trials for each condition.
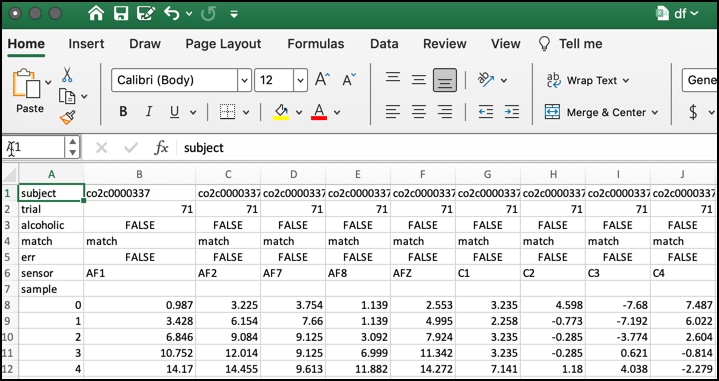
The .gz trial file is the object passed in the function import_eeg_file, contained in the EEG_load_function.py file. The function returns a dataframe with 64 columns, one for each electrode (a.k.a. channels).
FINAL OUTCOME FROM FULL DATASET
At the end, the script should run on the full dataset,eeg_full.tar, and return six dataframes for the means, and six dataframes for the standard deviation.
Each one of the six dataframes represents the average for a condition for a group. So for the means we will have:
- average frequency for control group, single stimulus, per time point per channel
- average frequency for control group, double stimulus matched, per time point per channel
- average frequency for control group, double stimulus non matched, per time point per channel
- average frequency for alcoholic group, single stimulus, per time point per channel
- average frequency for alcoholic group, double stimulus matched, per time point per channel
- average frequency for alcoholic group, double stimulus non matched, per time point per channel
Same for the variances.
Each one of the dataframes will have a header row and 256 rows, one for each timepoint. It will have one index column for the timepoints, and then 64 columns, one for each channel.
Basically, for what concerns the structure of the spreadsheet, each dataset looks like the output of import_eeg_file: time points as rows and channels as columns.
The only difference is that import_eeg_file's output header of a single .gz file looks like this:
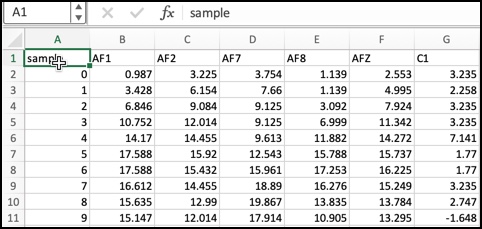
Instead the header of the dataframe with the averages will look lile this (obviously with different numbers):