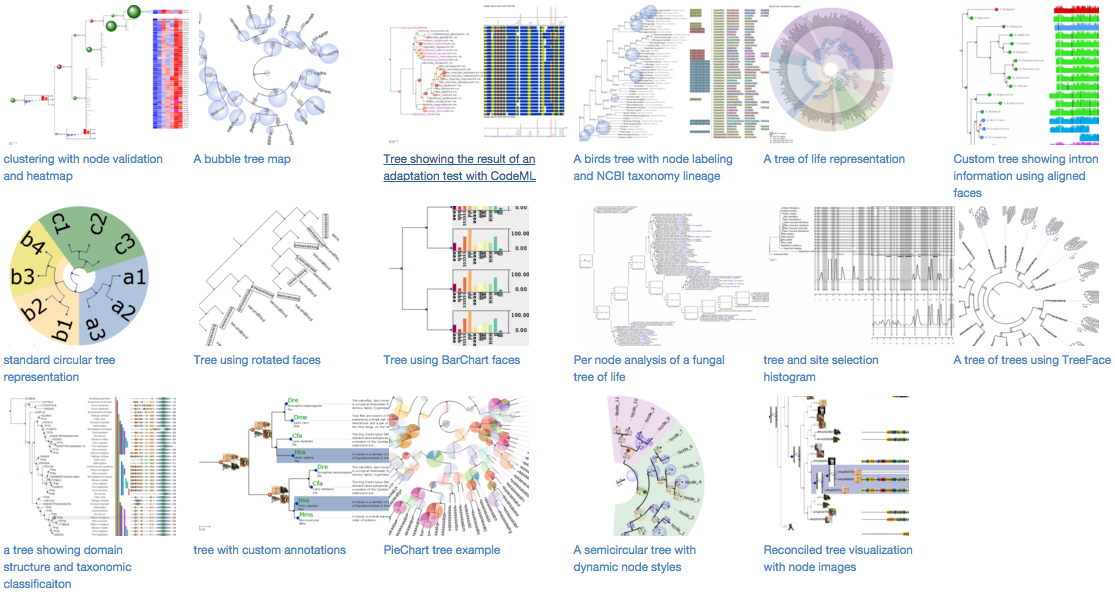
ETE (Environment for Tree Exploration) is a Python programming toolkit that assists in the automated manipulation, analysis and visualization of phylogenetic trees. Clustering trees or any other tree-like data structure are also supported.
Given that ETE is mainly developed as a tool for researchers working in phylogenetics and genomics, it also provides specialized tools in that context (e.g. reconstructing, comparing and visualizing phylogenetic trees). If you use ETE in a published work, please cite:
Jaime Huerta-Cepas, François Serra and Peer Bork. "ETE 3: Reconstruction, analysis and visualization of phylogenomic data." Mol Biol Evol (2016) doi: 10.1093/molbev/msw046
- The official web site of ETE is http://etetoolkit.org. Downloading instructions and further documentation can be found there.
- News and announcements are usually posted on twitter: http://twitter.com/etetoolkit
Please, whenerver possible, avoid sending direct support-related emails to the developers. Keep communication public:
For any type of question on how to use ETE in the bioinformatics context, use BioStars (http://biostars.org) or even StackOverflow forums.
Please use the "etetoolkit" tag for your questions:
Bug reports, feature requests and general discussion should be posted into github: https://github.com/etetoolkit/ete/issues
For more technical problems, you can also use the official ETE mailing list at https://groups.google.com/d/forum/etetoolkit. To avoid spam, messages from new users are moderated. Expect some delay until your first message and account is validated.
For any other inquire (collaborations, sponsoring, etc), please contact jhcepas /at/ gmail.com
https://github.com/etetoolkit/ete/wiki/Contributing