|
19 | 19 | "4. Compiling the generated code\n", |
20 | 20 | "5. Simulating the model\n", |
21 | 21 | "\n", |
22 | | - "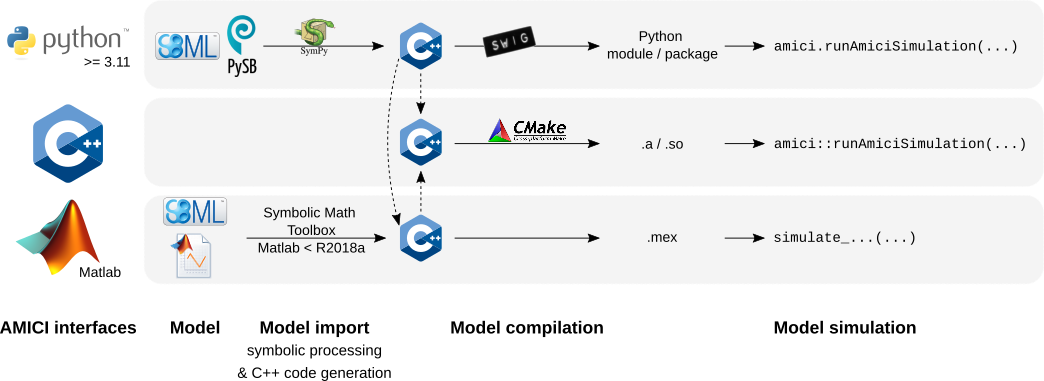\n", |
| 22 | + "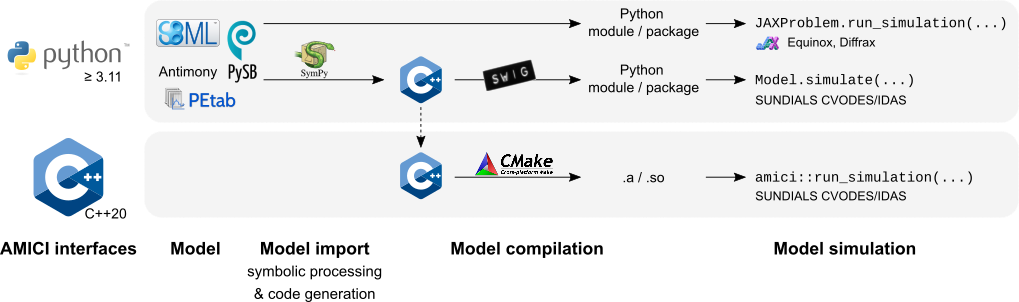\n", |
23 | 23 | "\n", |
24 | 24 | "There are various options to speed up individual steps of this process. Generally, faster import comes with slower simulation and vice versa. During parameter estimation, a model is often imported only once, and then millions of simulations are run. Therefore, faster simulation will easily compensate for slower import (one-off cost). In other cases, many models may to have to be imported, but only few simulations will be executed. In this case, faster import may be more relevant.\n", |
25 | 25 | "\n", |
|
71 | 71 | "\n", |
72 | 72 | "#### Fixing parameters\n", |
73 | 73 | "\n", |
74 | | - "By default, AMICI will generate sensitivity equations with respect to all model parameters. If it is clear upfront, that sensitivities with respect to certain parameters will not be required, their IDs can be passed to [amici.importers.sbml.SbmlImporter.sbml2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.sbml.SbmlImporter.html#amici.importers.sbml.SbmlImporter.sbml2amici) or [amici.importers.pysb.pysb2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.pysb.html?highlight=pysb2amici#amici.importers.pysb.pysb2amici) via the `fixed_parameters` argument to not generate the respective equations. This will reduce CPU time and RAM requirements during import and simulation.\n", |
| 74 | + "By default, AMICI will generate sensitivity equations with respect to all model parameters. If it is clear upfront, that sensitivities with respect to certain parameters will not be required, their IDs can be passed to [amici.importers.sbml.SbmlImporter.sbml2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.sbml.html#amici.importers.sbml.SbmlImporter.sbml2amici) or [amici.importers.pysb.pysb2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.pysb.html#amici.importers.pysb.pysb2amici) via the `fixed_parameters` argument to not generate the respective equations. This will reduce CPU time and RAM requirements during import and simulation.\n", |
75 | 75 | "The PEtab import will automatically pass all parameters with `petab.ESTIMATE==False` as `fixed_parameters` arguments.\n", |
76 | 76 | "\n", |
77 | 77 | "See also the following section for the case that no sensitivities are required at all.\n", |
78 | 78 | "\n", |
79 | 79 | "\n", |
80 | 80 | "#### Not generating sensitivity code\n", |
81 | 81 | "\n", |
82 | | - "If only forward simulations of a model are required, a modest import speedup can be obtained from not generating sensitivity code. This can be enabled via the `generate_sensitivity_code` argument of [amici.importers.sbml.SbmlImporter.sbml2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.sbml.SbmlImporter.html#amici.importers.sbml.SbmlImporter.sbml2amici) or [amici.importers.pysb.pysb2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.pysb.html?highlight=pysb2amici#amici.importers.pysb.pysb2amici).\n", |
| 82 | + "If only forward simulations of a model are required, a modest import speedup can be obtained from not generating sensitivity code. This can be enabled via the `generate_sensitivity_code` argument of [amici.importers.sbml.SbmlImporter.sbml2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.sbml.html#amici.importers.sbml.SbmlImporter.sbml2amici) or [amici.importers.pysb.pysb2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.pysb.html#amici.importers.pysb.pysb2amici).\n", |
83 | 83 | "\n", |
84 | 84 | "Example:\n", |
85 | 85 | "```bash\n", |
|
219 | 219 | "\n", |
220 | 220 | "By default, AMICI will try to perform some basic simplification of model expressions. For complex model expressions, or for large models, this can become costly. It very much depends on the model expressions, whether the benefits outweigh the cost.\n", |
221 | 221 | "\n", |
222 | | - "Simplification of model expressions can be disabled by passing `simplify=None` to [amici.importers.sbml.SbmlImporter.sbml2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.sbml.SbmlImporter.html#amici.importers.sbml.SbmlImporter.sbml2amici) or [amici.importers.pysb.pysb2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.pysb.html?highlight=pysb2amici#amici.importers.pysb.pysb2amici).\n", |
| 222 | + "Simplification of model expressions can be disabled by passing `simplify=None` to [amici.importers.sbml.SbmlImporter.sbml2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.sbml.html#amici.importers.sbml.SbmlImporter.sbml2amici) or [amici.importers.pysb.pysb2amici](https://amici.readthedocs.io/en/latest/generated/amici.importers.pysb.html#amici.importers.pysb.pysb2amici).\n", |
223 | 223 | "\n", |
224 | 224 | "Depending on the given model, different simplification schemes may be cheaper or more beneficial than the default. SymPy's simplification functions are [well documented](https://docs.sympy.org/latest/modules/simplify/simplify.html)." |
225 | 225 | ] |
|
477 | 477 | "\n", |
478 | 478 | "### Sensitivities w.r.t. a subset of parameters\n", |
479 | 479 | "\n", |
480 | | - "If only sensitivities with respect to a subset of model parameters are of interest to you (see also *Fixing parameters* above), you can speed up the simulation by selecting the relevant parameter indices via [amici.Model.setParameterList](https://amici.readthedocs.io/en/latest/generated/amici.amici.Model.html#amici.amici.Model.setParameterList).\n" |
| 480 | + "If only sensitivities with respect to a subset of model parameters are of interest to you (see also *Fixing parameters* above), you can speed up the simulation by selecting the relevant parameter indices via [amici.sim.sundials.Model.set_parameter_list](https://amici.readthedocs.io/en/latest/generated/amici.sim.sundials.html#amici.sim.sundials.Model.set_parameter_list).\n" |
481 | 481 | ] |
482 | 482 | }, |
483 | 483 | { |
484 | 484 | "cell_type": "markdown", |
485 | 485 | "id": "af4bd3d5", |
486 | 486 | "metadata": {}, |
487 | 487 | "source": [ |
488 | | - "\n", |
489 | 488 | "### Parallel simulation of multiple conditions\n", |
490 | 489 | "\n", |
491 | | - "Whenever there are multiple independent simulations to perform, you can use [amici.runAmiciSimulations(..., num_threads=...)](https://amici.readthedocs.io/en/latest/generated/amici.amici.html#amici.amici.runAmiciSimulations) instead of [amici.runAmiciSimulations(...)](https://amici.readthedocs.io/en/latest/generated/amici.amici.html#amici.amici.runAmiciSimulation) to run them in parallel. Note that all simulation results have to be kept in memory, which may become problematic for very large numbers of simulations.\n", |
| 490 | + "Whenever there are multiple independent simulations to perform, you can use [amici.sim.sundials.run_simulations(..., num_threads=...)](https://amici.readthedocs.io/en/latest/generated/amici.sim.sundials.html#amici.sim.sundials.run_simulations) instead of `amici.sim.sundials.run_simulations(...)` to run them in parallel. Note that all simulation results have to be kept in memory, which may become problematic for very large numbers of simulations.\n", |
492 | 491 | "Parallelization is based on OpenMP and does not come with the issues associated with Python's multiprocessing or multithreading (spawning extra processes or limitations related to the global interpreter lock).\n", |
493 | 492 | "\n", |
494 | 493 | "### Reporting mode\n", |
495 | 494 | "\n", |
496 | 495 | "During model simulation, many quantities are calculated, but not all might be of interest for you. For example, for parameter estimation you might only be interested in the likelihood and gradient. In this case, you can save time and memory using\n", |
497 | | - "[amici.Solver.setReturnDataReportingMode(amici.RDataReporting.likelihood)](https://amici.readthedocs.io/en/latest/generated/amici.amici.Solver.html#amici.amici.Solver.setReturnDataReportingMode)." |
| 496 | + "[amici.sim.sundials.Solver.set_return_data_reporting_mode(amici.sim.sundials.RDataReporting.likelihood)](https://amici.readthedocs.io/en/latest/generated/amici.sim.sundials.html#amici.sim.sundials.Solver.set_return_data_reporting_mode)." |
498 | 497 | ] |
499 | 498 | } |
500 | 499 | ], |
|
0 commit comments